Liana Barkan MD
Faculty Peer ReviewedÂ
So much mystery and confusion, and yet so few answers, surround the current H1N1 Pandemic. From where did it come? How did it evolve to have genes from avian, human, and swine flu viruses? How does a virus that normally requires direct contact with the source animal develop the ability to sustain human-to-human transmission? What determines its mechanism of pathogenicity? Before we can attempt to answer these questions we need to review the basic pathophysiology of the influenza virus and its transmission.
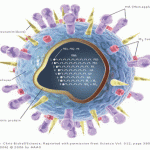
The influenza virus belongs to the family of Orthomyxoviridae viruses, in which influenza types A and B belong to one genus and type C belongs to another. Antigenic characteristics of the nucleoprotein antigens and the matrix antigens determine the designation. Influenza viruses are also subdivided based on the properties of two types of surface glycoproteins projecting from the lipid envelope, known as the hemagglutinin (H) and neuraminidase (N) antigens (see Figure-1). Hemagglutinin binds to host cell receptors and neuraminidase subsequently degrades the cell receptor. Following viral replication, neuraminidase may also play a roll in releasing the viral progeny from the infected cell.
Influenza virus normally causes a predominant respiratory illness accompanied by non-specific constitutional symptoms, such as fevers, chills, headache, myalgias, and general malaise. Antibodies we form against the hemagglutinin antigen confer immunity to the influenza virus, whereas our antibodies against neuraminidase limit viral spread and severity of infection. However, despite vaccination efforts, the influenza A virus persists in the population. This is likely due to the reassortment of its eight single-stranded RNA segments during an infection and particularly in situations in which a cell becomes infected with more than one type of influenza A virus.[1] In addition to reassortment, antigenic mutations in the hemagglutinin and neuraminidase proteins lead to either antigenic shifts (major variations) or antigenic drifts (minor variations) that are responsible for some of the more severe outbreaks and pandemics seen since the early 1900’s.1
Figure 1
In an article published in May 2009 in the New England Journal of Medicine, Shinde et al. suggest that pigs might act as mixing vessels for the triple reassortment of avian, swine, and human influenza viruses and may have caused sporadic cases of infection in humans as early as the late 1990s, well before the current Swine Flu Epidemic.[2] In fact, a swine influenza virus causing febrile respiratory illness in pigs is well known to be enzootic to pigs in North America.[2] The sporadic cases in humans occurred in high-risk individuals working in direct contact with pig herds and to a much lesser extent those in contact with the people who worked with pigs. However, until the recent outbreak that started in Mexico, little evidence existed for sustained human-to-human transmission of swine influenza virus.
Transmission of the influenza virus usually occurs via aerosols generated during a cough or a sneeze, but can also occur through other types of personal contact, and even possibly through intermediary objects, such as door knobs, desks, tables, or phones. In the article cited above, Shinde et al. found evidence for only limited human-to-human transmission of swine influenza virus. Of the 11 human cases with documented triple reassortment swine influenza, five had had direct contact with a pig; three had been within six feet of a pig; one had been in the near vicinity; one had had contact with someone with a suspected case of swine influenza; and one had had no known contact with a pig. In contrast, one novel aspect of the current pandemic is that the majority of cases have not been linked to exposure to pigs and instead appear to demonstrate sustained human-to-human transmission.
In comparison to previous outbreaks, this current virus also demonstrates much more dramatically efficient transmission among people as evidenced by its rapid and prolific spread. According to a recent CDC update, although the current virus was originally referred to as “swine flu” due to genetic similarity to the enzootic influenza virus occurring in pigs, further study has shown that the new virus is actually very different. Swine flu typically only affects people in direct contact with pigs, whereas the human influenza virus spreads predominately from person to person. The new current virus has been called a “quadruple reassortment” virus because it contains swine genes from both Europe and Asia, avian genes, and human genes.[3] Shinde et al. revealed that in 10 of the 11 isolated influenza viruses in their study, five of the eight single-stranded RNA gene segments were of North American swine origin, two were of North American avian origin, and one was of human origin. Work is still being done to further classify the genomic sequences in the current outbreak and may help us better understand why the mode of transmission and its sustainability of the current virus appear to be so different from previous strains.
Unfortunately at this time, there are still more questions than answers, but the take home message remains the same: cover your mouth when you cough or sneeze, wash your hands frequently with antibacterial soaps or alcohol based cleansers, avoid close contact with sick people, and stay home if you are sick.
As of July 12 2009 at 11:00 AM ET, the CDC reports 37,246 confirmed and probable cases and 211 deaths in the United States. 3 WHO reports 94512 Cases with 429 deaths worldwide. [4]
Dr. Barkan is a 3rd year resident in internal medicine at NYU Medical Center.
Reviewed by Howard Leaf, MD, Assistant Professor, NYU Division of Infectious Diseases and ImmunologyÂ
References:
1. Fauci AS, et. al, Harrison’s Principles of Internal Medicine, 14th ed, McGraw-Hill, 1998, pp 1112-1116
2. Shinde V, et al. “Triple-Reassortant Swine Influenza A (H1) in Humans in the United States, 2005-2009, NEJM.org, May 7th, 2009
3. http://www.cdc.gov/h1n1flu/qa.htm.
4. http://www.who.int/csr/don/2009_07_06/en/index.html